
AI CERTS
2 hours ago
CellSAM: AI Biotech Breakthrough For Cell Imaging
Moreover, early adopters report dramatic reductions in annotation time. This introduction sets the stage for a detailed look at how CellSAM alters laboratory routines and corporate pipelines.
However, excitement must be balanced with rigor. Therefore, this article dissects design choices, training logistics, datasets, benchmarks, and practical deployment advice. Each section ends with concise takeaways, ensuring busy professionals extract value quickly. Throughout, the term AI Biotech appears as a thematic anchor, highlighting the field’s rapid convergence of computation and biology.
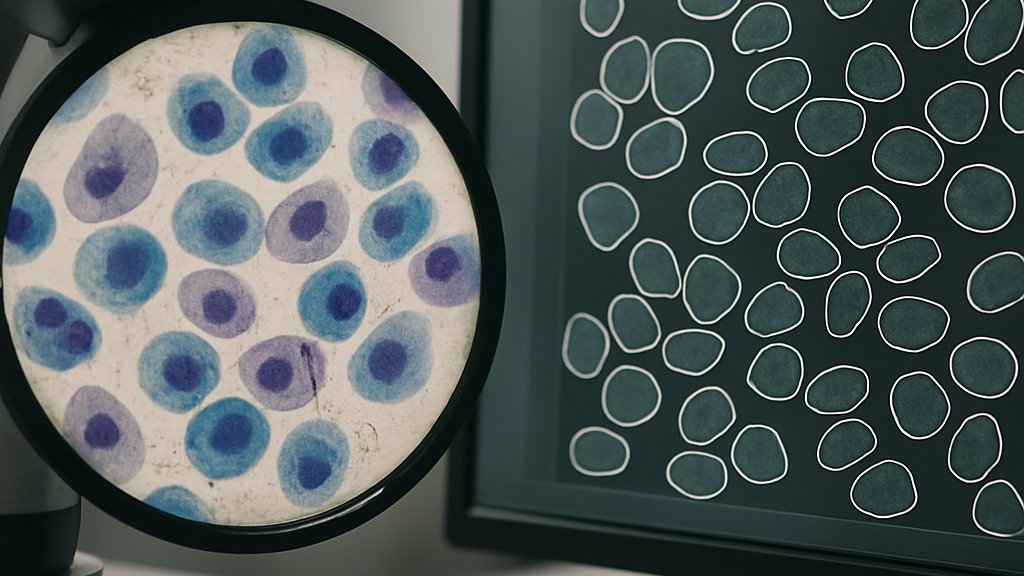
CellSAM Model Design Details
CellSAM builds on SAM’s Vision Transformer backbone. Additionally, the Caltech team added an object detector named CellFinder. This module uses Anchor DETR to generate bounding boxes automatically. Subsequently, the boxes feed into SAM’s promptable decoder to yield crisp instance masks. The approach differs from pixel-dense networks like Cellpose. Instead, CellSAM treats every cell as a separate query. In contrast, earlier models processed entire images in one sweep.
Developers fine-tuned only SAM’s neck layers, keeping its rich encoder frozen. Consequently, training remained stable despite limited biological data. Furthermore, prompt engineering became trivial because CellFinder handled bounding-box generation at scale. AI Biotech startups value this modular blueprint because components can evolve independently.
These architectural insights emphasize flexibility. However, implementation means nothing without robust training, which we address next.
Training Workflow And Compute
The CellFinder detector trained for 2,800 epochs. Batch size stayed at four due to GPU memory limits. Training consumed six days and twelve hours on eight A100 GPUs. Meanwhile, final fine-tuning of the composite system required only one day on a single RTX 4090. Therefore, small academic groups can reproduce results without massive clusters.
Energy efficiency also matters. Moreover, running one SAM query per cell increases inference cost on dense slides. Nevertheless, the modular setup allows batched processing on modern consumer cards. Many Biotech companies already own compatible hardware.
AI Biotech teams should budget compute wisely. Yet hardware is useless without diverse data. The next section covers that critical ingredient.
Dataset Breadth And Diversity
Caltech aggregated images from roughly ten public and internal sources. These included TissueNet, DeepBacs, BriFiSeg, OmniPose, YeastNet, and Kaggle’s DSB. Additionally, H&E pathology slides and Phase-contrast sets expanded coverage. Sample counts spanned mammalian, bacterial, and yeast taxa.
The authors deliberately held out LIVECell for zero-shot evaluation. Consequently, they avoided data leakage and measured honest generalization. Such curation demonstrates mature experimental discipline often missing in emergent AI Biotech projects.
- 330 tissue fields
- 144 cell-culture fields
- 51 H&E fields
- 260 bacterial images
- 32 yeast images
- 56 nuclear channels
The range above underpins strong transfer ability. These numbers also justify future cross-lab collaborations. However, metrics finally decide real-world value, as explained next.
Benchmark Results And Impact
Performance used the inverse F1 score. Lower values indicate better masks. CellSAM cut error across every modality compared with Cellpose. On the challenging LIVECell zero-shot test, the model lifted F1 from 0.13 to 0.40. Few-shot adaptation with ten fields often reached human agreement. Furthermore, inter-rater studies showed parity between CellSAM and expert annotators.
Key implications appear below.
- Human-level accuracy across fluorescence, brightfield, and H&E
- Box prompting reduced annotation clicks by an order of magnitude
- Few-shot tuning requires roughly one afternoon per new cell line
Consequently, pharmaceutical screening teams can track millions of cells over time. Moreover, spatial biology startups gain reliable masks for downstream quantification. These outcomes mark a milestone for AI Biotech research. Nevertheless, successful lab adoption needs workflow guidance, which follows.
Practical Lab Adoption Guide
First, scientists should evaluate baseline zero-shot accuracy on their own images. Subsequently, collect ten representative fields and perform few-shot fine-tuning. DeepCell Label integrates the process within a web browser. Furthermore, bounding-box annotation feels natural even for novices.
Professionals can enhance expertise with the AI Researcher™ certification. The program covers promptable segmentation, transfer learning, and regulatory considerations. Consequently, graduates quickly align laboratory needs with production-grade AI Biotech pipelines.
After training, schedule inference jobs during off-peak hours. Meanwhile, monitor GPU memory because each mask request spawns a new forward pass. In contrast, dense models saturate memory once per image. Therefore, throughput decisions depend on cell density.
These steps streamline deployment. However, limitations still exist, demanding careful attention next.
Limitations And Future Work
Cell morphology far from the training distribution can hinder accuracy. For example, SH-SY5Y neuroblastoma cells challenge the detector. Moreover, bounding-box prompts struggle with filamentous bacteria. Therefore, supplementary manual correction remains necessary.
Inference speed declines for slides containing thousands of tightly packed cells. Consequently, high-throughput core facilities might prefer hybrid pipelines that combine CellSAM with pixel-dense coarse passes. Additionally, dataset bias persists despite thoughtful curation. Independent validation on proprietary datasets remains essential for any Biotech venture.
Future research should benchmark whole-slide pathology throughput. Moreover, third-party labs must publish replication studies in Nature or other journals. These efforts will cement trust and reveal improvement avenues.
The challenges noted above inform next steps. However, a broader summary will help decision makers act decisively.
Conclusion And Next Steps
CellSAM delivers a versatile segmentation foundation model. It merges SAM’s flexibility with a transformer detector, offering strong zero-shot performance. Moreover, few-shot adaptation unlocks near human quality within hours. The system trims annotation workloads dramatically and empowers large-scale Imaging studies.
Nevertheless, compute costs and edge-case morphologies require ongoing attention. Consequently, labs should validate performance locally and iterate. Professionals seeking deeper mastery can pursue the linked certification and position themselves at the forefront of AI Biotech innovation.
Adopt CellSAM today, experiment diligently, and share findings. The coming year will reward teams who bridge biological insight with robust computational strategy.
Disclaimer: Some content may be AI-generated or assisted and is provided ‘as is’ for informational purposes only, without warranties of accuracy or completeness, and does not imply endorsement or affiliation.



